Feature-Klassifikation verbessere met projizierte Quantekerne
Zosamme jerechnet: 80 Minute op enem Heron r3 Prozessor (HENJANG: Dat es blooß ene Schätzwärt. Ding Zick kann angers sinn.)
En dämm Tutorial zeije mer, wi mer ene projected quantum kernel (PQK) met Qiskit op enem echte biolojesche Datensatz loufe losse kann, opjebaue op däm Paper Enhanced Prediction of CAR T-Cell Cytotoxicity with Quantum-Kernel Methods [1].
PQK es en Methood, di mer em quantum machine learning (QML) bruch, öm klassische Date en ene Quante-Feature-Ruum zo kodiere un se dann widder retuur en de klassische Domäne ze projiziere, endämm mer Quantecomputer för verbesserdi Feature-Uswahl nutz. Dat bestoht doruß, klassische Date en Quantezöständ ze kodiere met enem Quanteschaltkreis, typischerwiis dörch e Prozess di mer Feature Mapping nenne, wo di Date en ene höchdimensionale Hilbert-Ruum ömjewandelt wäde. Dä "projected" Aspek bezügg sich drop, klassische Informatione uß dä Quantezöständ ze extrahiere, endämm mer bestemmpte Observabele messe, öm en Kernel-Matrix ze baue, di mer en klassische Kernel-basierde Algorithme wi Support Vector Machines verwende kann. Dä Ansatz nutz di rechnerische Vördeile vun Quantesysteme, öm möjlicherwiis besser Leistung för bestemmpte Ufjaabe em Verjlich met klassische Methode ze erziehle.
Dat Tutorial nimmt och aan, dat mer allgemein met QML-Methode vertraut sinn. För wigger Erkundung vun QML, luurt noh däm Quantum machine learning Kurs em IBM Quantum Learning.
Aanforderunge
Bevoer mer met dämm Tutorial aanfange, stellt secher, dat Ühr dat folgende installiert habt:
- Qiskit SDK v2.0 oder späder, met visualization Ungerstötzung
- Qiskit Runtime v0.40 oder späder (
pip install qiskit-ibm-runtime) - Category encoders 2.8.1 (
pip install category-encoders) - NumPy 2.3.2 (
pip install numpy) - Pandas 2.3.2 (
pip install pandas) - Scikit-learn 1.7.1 (
pip install scikit-learn) - Tqdm 4.67.1 (
pip install tqdm)
Setup
# Added by doQumentation — required packages for this notebook
!pip install -q category-encoders numpy pandas qiskit qiskit-ibm-runtime scipy scikit-learn tqdm
import warnings
# Standard libraries
import os
import numpy as np
import pandas as pd
# Machine learning and data processing
import category_encoders as ce
from scipy.linalg import inv, sqrtm
from sklearn.metrics.pairwise import rbf_kernel
from sklearn.model_selection import GridSearchCV, StratifiedKFold
from sklearn.svm import SVC
# Qiskit and Qiskit Runtime
from qiskit import QuantumCircuit
from qiskit.circuit import ParameterVector
from qiskit.circuit.library import UnitaryGate, ZZFeatureMap
from qiskit.quantum_info import SparsePauliOp, random_unitary
from qiskit.transpiler import generate_preset_pass_manager
from qiskit_ibm_runtime import (
Batch,
EstimatorOptions,
EstimatorV2 as Estimator,
QiskitRuntimeService,
)
# Progress bar
import tqdm
warnings.filterwarnings("ignore")
Schrett 1: Klassische Injaabe en e Quanteproblem mappe
Datensatz vörbereide
En dämm Tutorial brauche mer ene echte biolojesche Datensatz för en binär Klassifikationsufjaab, dä vun Daniels et al. (2022) jeneriert wood un uß däm supplementary material eronjerjelade wäde kann, dat met däm Paper medjelivvert wood. Di Date bestonn uß CAR T-Zelle, di sin jenetisch veränderte T-Zelle, di mer en de Immuntherapie för di Behandlung vun bestemmpte Krebs bruch. T-Zelle, en Art vun Immunzell, wäde em Labor ömjestalt, öm chimär Antigene-Rezeptore (CARs) auszudrücke, di op bestemmpte Proteine op Krebszelle ziele. Diss veränderte T-Zelle künne Krebszelle effektiver erkenne un zerstöre. Di Date-Features sin di CAR T-Zell Motive, di sich op di bestemmpte strukturäle oder funktionäle Komponente vun däm CAR beziehe, dä en T-Zelle enjebout wood. Opjebaue op disse Motive, es uns Ufjaab, di Cytotoxizität vun ener jejävene CAR T-Zell vörherzesare, un se als entweder toxisch oder nit-toxisch ze kennzeichne. Dat folgende zeich di Hëlfsfunktione, öm dämm Datensatz vörzeverarbeide.
def preprocess_data(dir_root, args):
"""
Preprocess the training and test data.
"""
# Read from the csv files
train_data = pd.read_csv(
os.path.join(dir_root, args["file_train_data"]),
encoding="unicode_escape",
sep=",",
)
test_data = pd.read_csv(
os.path.join(dir_root, args["file_test_data"]),
encoding="unicode_escape",
sep=",",
)
# Fix the last motif ID
train_data[train_data == 17] = 14
train_data.columns = [
"Cell Number",
"motif",
"motif.1",
"motif.2",
"motif.3",
"motif.4",
"Nalm 6 Cytotoxicity",
]
test_data[test_data == 17] = 14
test_data.columns = [
"Cell Number",
"motif",
"motif.1",
"motif.2",
"motif.3",
"motif.4",
"Nalm 6 Cytotoxicity",
]
# Adjust motif at the third position
if args["filter_for_spacer_motif_third_position"]:
train_data = train_data[
(train_data["motif.2"] == 14) | (train_data["motif.2"] == 0)
]
test_data = test_data[
(test_data["motif.2"] == 14) | (test_data["motif.2"] == 0)
]
train_data = train_data[
args["motifs_to_use"] + [args["label_name"], "Cell Number"]
]
test_data = test_data[
args["motifs_to_use"] + [args["label_name"], "Cell Number"]
]
# Adjust motif at the last position
if not args["allow_spacer_motif_last_position"]:
last_motif = args["motifs_to_use"][len(args["motifs_to_use"]) - 1]
train_data = train_data[
(train_data[last_motif] != 14) & (train_data[last_motif] != 0)
]
test_data = test_data[
(test_data[last_motif] != 14) & (test_data[last_motif] != 0)
]
# Get the labels
train_labels = np.array(train_data[args["label_name"]])
test_labels = np.array(test_data[args["label_name"]])
# For the classification task use the threshold to binarize labels
train_labels[train_labels > args["label_binarization_threshold"]] = 1
train_labels[train_labels < 1] = args["min_label_value"]
test_labels[test_labels > args["label_binarization_threshold"]] = 1
test_labels[test_labels < 1] = args["min_label_value"]
# Reduce data to just the motifs of interest
train_data = train_data[args["motifs_to_use"]]
test_data = test_data[args["motifs_to_use"]]
# Get the class and motif counts
min_class = np.min(np.unique(np.concatenate([train_data, test_data])))
max_class = np.max(np.unique(np.concatenate([train_data, test_data])))
num_class = max_class - min_class + 1
num_motifs = len(args["motifs_to_use"])
print(str(max_class) + ":" + str(min_class) + ":" + str(num_class))
train_data = train_data - min_class
test_data = test_data - min_class
return (
train_data,
test_data,
train_labels,
test_labels,
num_class,
num_motifs,
)
def data_encoder(args, train_data, test_data, num_class, num_motifs):
"""
Use one-hot or binary encoding for classical data representation.
"""
if args["encoder"] == "one-hot":
# Transform to one-hot encoding
train_data = np.eye(num_class)[train_data]
test_data = np.eye(num_class)[test_data]
train_data = train_data.reshape(
train_data.shape[0], train_data.shape[1] * train_data.shape[2]
)
test_data = test_data.reshape(
test_data.shape[0], test_data.shape[1] * test_data.shape[2]
)
elif args["encoder"] == "binary":
# Transform to binary encoding
encoder = ce.BinaryEncoder()
base_array = np.unique(np.concatenate([train_data, test_data]))
base = pd.DataFrame(base_array).astype("category")
base.columns = ["motif"]
for motif_name in args["motifs_to_use"][1:]:
base[motif_name] = base.loc[:, "motif"]
encoder.fit(base)
train_data = encoder.transform(train_data.astype("category"))
test_data = encoder.transform(test_data.astype("category"))
train_data = np.reshape(
train_data.values, (train_data.shape[0], num_motifs, -1)
)
test_data = np.reshape(
test_data.values, (test_data.shape[0], num_motifs, -1)
)
train_data = train_data.reshape(
train_data.shape[0], train_data.shape[1] * train_data.shape[2]
)
test_data = test_data.reshape(
test_data.shape[0], test_data.shape[1] * test_data.shape[2]
)
else:
raise ValueError("Invalid encoding type.")
return train_data, test_data
Ühr künnt dat Tutorial loufe losse, endämm mer di folgende Zell usföhrt, di automatisch di nüdije Ordner-Struktur erstellt un söwohl di Trainings- als och di Test-Dateie direkt en Ühr Ömjevung eronjerlaadt. Wann Ühr diss Dateie schonn lokal habt, weed dämm Schrett se secher överschriive, öm Version-Konsistenz ze jewohrleiste.
## Download dataset
# Create data directory if it doesn't exist
!mkdir -p data_tutorial/pqk
# Download the training and test sets from the official Qiskit documentation repo
!wget -q --show-progress -O data_tutorial/pqk/train_data.csv \
https://raw.githubusercontent.com/Qiskit/documentation/main/datasets/tutorials/pqk/train_data.csv
!wget -q --show-progress -O data_tutorial/pqk/test_data.csv \
https://raw.githubusercontent.com/Qiskit/documentation/main/datasets/tutorials/pqk/test_data.csv
!wget -q --show-progress -O data_tutorial/pqk/projections_train.csv \
https://raw.githubusercontent.com/Qiskit/documentation/main/datasets/tutorials/pqk/projections_train.csv
!wget -q --show-progress -O data_tutorial/pqk/projections_test.csv \
https://raw.githubusercontent.com/Qiskit/documentation/main/datasets/tutorials/pqk/projections_test.csv
# Check the files have been downloaded
!echo "Dataset files downloaded:"
!ls -lh data_tutorial/pqk/*.csv
args = {
"file_train_data": "train_data.csv",
"file_test_data": "test_data.csv",
"motifs_to_use": ["motif", "motif.1", "motif.2", "motif.3"],
"label_name": "Nalm 6 Cytotoxicity",
"label_binarization_threshold": 0.62,
"filter_for_spacer_motif_third_position": False,
"allow_spacer_motif_last_position": True,
"min_label_value": -1,
"encoder": "one-hot",
}
dir_root = "./"
# Preprocess data
train_data, test_data, train_labels, test_labels, num_class, num_motifs = (
preprocess_data(dir_root=dir_root, args=args)
)
# Encode the data
train_data, test_data = data_encoder(
args, train_data, test_data, num_class, num_motifs
)
14:0:15
Mer transformiere däm Datensatz och esu, dat als för Skalierungszwecke darjestellt weed.
# Change 1 to pi/2
angle = np.pi / 2
tmp = pd.DataFrame(train_data).astype("float64")
tmp[tmp == 1] = angle
train_data = tmp.values
tmp = pd.DataFrame(test_data).astype("float64")
tmp[tmp == 1] = angle
test_data = tmp.values
Mer prööve di Jrößte un Forme vun dä Trainings- un Test-Datensätze.
print(train_data.shape, train_labels.shape)
print(test_data.shape, test_labels.shape)
(172, 60) (172,)
(74, 60) (74,)
Schrett 2: Problem för Quante-Hardware-Usföhrung optimiere
Quanteschaltkreis
Mer baue jetz di Feature Map, di uns klassische Datensatz en ene höherdimensionale Feature-Ruum embettet. För diss Embedding brauche mer dat ZZFeatureMap uß Qiskit.
feature_dimension = train_data.shape[1]
reps = 24
insert_barriers = True
entanglement = "pairwise"
# ZZFeatureMap with linear entanglement and a repetition of 2
embed = ZZFeatureMap(
feature_dimension=feature_dimension,
reps=reps,
entanglement=entanglement,
insert_barriers=insert_barriers,
name="ZZFeatureMap",
)
embed.decompose().draw(output="mpl", style="iqp", fold=-1)

En anger Quante-Embedding-Option es dä 1D-Heisenberg Hamiltonian Evolution Ansatz. Ühr künnt dat Afschnett övverspringe, wann Ühr met däm ZZFeatureMap wigger maache wollt.
feature_dimension = train_data.shape[1]
num_qubits = feature_dimension + 1
embed2 = QuantumCircuit(num_qubits)
num_trotter_steps = 6
pv_length = feature_dimension * num_trotter_steps
pv = ParameterVector("theta", pv_length)
# Add Haar random single qubit unitary to each qubit as initial state
np.random.seed(42)
seeds_unitary = np.random.randint(0, 100, num_qubits)
for i in range(num_qubits):
rand_gate = UnitaryGate(random_unitary(2, seed=seeds_unitary[i]))
embed2.append(rand_gate, [i])
def trotter_circ(feature_dimension, num_trotter_steps):
num_qubits = feature_dimension + 1
circ = QuantumCircuit(num_qubits)
# Even
for i in range(0, feature_dimension, 2):
circ.rzz(2 * pv[i] / num_trotter_steps, i, i + 1)
for i in range(0, feature_dimension, 2):
circ.rxx(2 * pv[i] / num_trotter_steps, i, i + 1)
for i in range(0, feature_dimension, 2):
circ.ryy(2 * pv[i] / num_trotter_steps, i, i + 1)
# Odd
for i in range(1, feature_dimension, 2):
circ.rzz(2 * pv[i] / num_trotter_steps, i, i + 1)
for i in range(1, feature_dimension, 2):
circ.rxx(2 * pv[i] / num_trotter_steps, i, i + 1)
for i in range(1, feature_dimension, 2):
circ.ryy(2 * pv[i] / num_trotter_steps, i, i + 1)
return circ
# Hamiltonian evolution ansatz
for step in range(num_trotter_steps):
circ = trotter_circ(feature_dimension, num_trotter_steps)
if step % 2 == 0:
embed2 = embed2.compose(circ)
else:
reverse_circ = circ.reverse_ops()
embed2 = embed2.compose(reverse_circ)
embed2.draw(output="mpl", style="iqp", fold=-1)
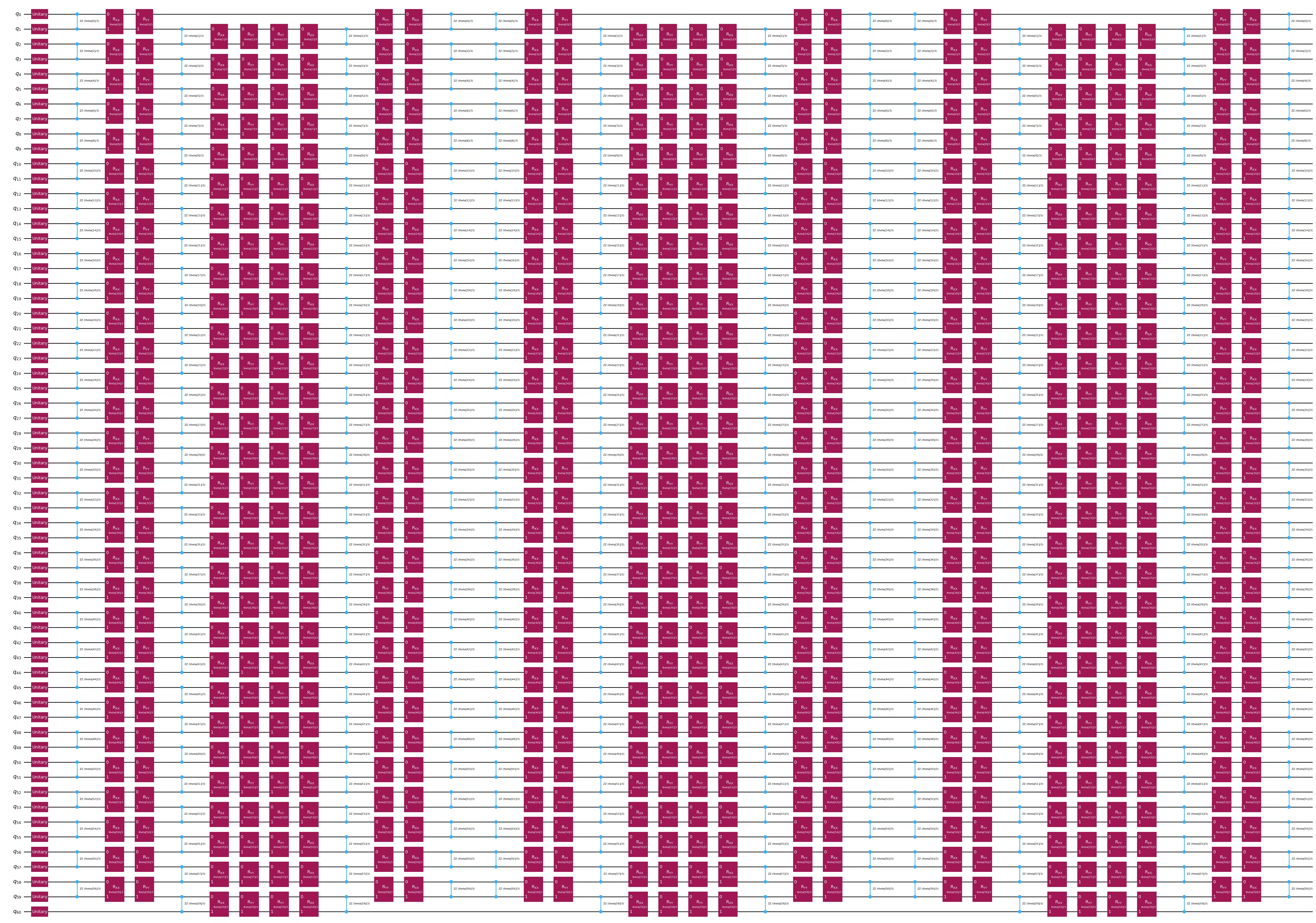
Schrett 3: Met Qiskit-Primitives usföhre
1-RDMs messe
Di wichtigste Bausteins vun projected quantum kernels sin di reduzierte Dichte-Matrixe (RDMs), di dörch projektive Messunge vun de Quante-Feature Map jekrääje wäde. En dämm Schrett kräje mer all Single-Qubit-reduzierte Dichte-Matrixe (1-RDMs), di später en di klassische exponentielle Kernel-Funktion jejevve wäde.
Loore mer uns aan, wi mer di 1-RDM för en einzelne Datepungk uß däm Datensatz berechne, bevoer mer över all Date loufe. Di 1-RDMs sin en Sammlung vun Single-Qubit-Messunge vun Pauli X, Y un Z Operatöre op all Qubits. Dat es esu, weil en Single-Qubit-RDM vollständig als usjedrückt wäde kann.
Zeesch wähle mer dat Backend uß, dat mer bruche welle.
service = QiskitRuntimeService()
backend = service.least_busy(
operational=True, simulator=False, min_num_qubits=133
)
target = backend.target
Dann loufe mer däm Quanteschaltkreis un messe di Projektione. Luurt druf, dat mer Fehlerminderung aaschalle, inklusiv Zero Noise Extrapolation (ZNE).
# Let's select the ZZFeatureMap embedding for this example
qc = embed
num_qubits = feature_dimension
# Identity operator on all qubits
id = "I" * num_qubits
# Let's select the first training datapoint as an example
parameters = train_data[0]
# Bind parameter to the circuit and simplify it
qc_bound = qc.assign_parameters(parameters)
transpiler = generate_preset_pass_manager(
optimization_level=3, basis_gates=["u3", "cz"]
)
transpiled_circuit = transpiler.run(qc_bound)
# Transpile for hardware
transpiler = generate_preset_pass_manager(optimization_level=3, target=target)
transpiled_circuit = transpiler.run(transpiled_circuit)
# We group all commuting observables
# These groups are the Pauli X, Y and Z operators on individual qubits
observables_x = [
SparsePauliOp(id[:i] + "X" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_y = [
SparsePauliOp(id[:i] + "Y" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_z = [
SparsePauliOp(id[:i] + "Z" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
# We define the primitive unified blocs (PUBs) consisting of the embedding circuit,
# set of observables and the circuit parameters
pub_x = (transpiled_circuit, observables_x)
pub_y = (transpiled_circuit, observables_y)
pub_z = (transpiled_circuit, observables_z)
# Experiment options for error mitigation
num_randomizations = 300
shots_per_randomization = 100
noise_factors = [1, 3, 5]
experimental_opts = {}
experimental_opts["resilience"] = {
"measure_mitigation": True,
"zne_mitigation": True,
"zne": {
"noise_factors": noise_factors,
"amplifier": "gate_folding",
"extrapolated_noise_factors": [0] + noise_factors,
},
}
experimental_opts["twirling"] = {
"num_randomizations": num_randomizations,
"shots_per_randomization": shots_per_randomization,
"strategy": "active-accum",
}
# We define and run the estimator to obtain <X>, <Y> and <Z> on all qubits
estimator = Estimator(mode=backend, options=experimental_opts)
job = estimator.run([pub_x, pub_y, pub_z])
Als nächstes holle mer di Erjebnisse.
job_result_x = job.result()[0].data.evs
job_result_y = job.result()[1].data.evs
job_result_z = job.result()[2].data.evs
print(job_result_x)
print(job_result_y)
print(job_result_z)
[ 3.67865951e-03 1.01158571e-02 -3.95790878e-02 6.33984326e-03
1.86035759e-02 -2.91533268e-02 -1.06374793e-01 4.48873518e-18
4.70201764e-02 3.53997968e-02 2.53130819e-02 3.23903401e-02
6.06327843e-03 1.16313667e-02 -1.12387504e-02 -3.18457725e-02
-4.16445718e-04 -1.45609602e-03 -4.21737114e-01 2.83705669e-02
6.91332890e-03 -7.45363001e-02 -1.20139326e-02 -8.85566135e-02
-3.22648394e-02 -3.24228074e-02 6.20431299e-04 3.04225434e-03
5.72795792e-03 1.11288428e-02 1.50395861e-01 9.18380197e-02
1.02553163e-01 2.98312847e-02 -3.30298912e-01 -1.13979648e-01
4.49159340e-03 8.63861493e-02 3.05666566e-02 2.21463145e-04
1.45946735e-02 8.54537275e-03 -8.09805979e-02 -2.92608104e-02
-3.91243644e-02 -3.96632760e-02 -1.41187613e-01 -1.07363243e-01
1.81089440e-02 2.70778895e-02 1.45139414e-02 2.99480458e-02
4.99137134e-02 7.08789852e-02 4.30565759e-02 8.71287156e-02
1.04334798e-01 7.72191962e-02 7.10059720e-02 1.04650403e-01]
[-7.31765102e-05 7.42669174e-03 9.82277344e-03 5.92638249e-02
4.24120486e-02 -9.06473416e-03 4.55057675e-03 8.43494094e-03
6.92097339e-02 -6.82234424e-02 6.13509008e-02 3.94200491e-02
-1.24037979e-02 1.01976642e-01 7.90538600e-03 -7.19726160e-02
-1.19501703e-16 -1.03796614e-02 7.37382463e-02 1.97238568e-01
-3.59250635e-02 -2.67554009e-02 3.55010633e-02 7.68877990e-02
6.50677589e-05 -6.59298767e-03 -1.23719487e-02 -6.41938151e-02
1.95603072e-02 -2.48448551e-02 5.17784810e-02 -5.93767100e-02
3.11897681e-02 -3.91959720e-18 -4.47769148e-03 1.39202197e-01
-6.56387523e-02 -5.85665483e-02 9.52905894e-03 -8.61460731e-02
3.91790656e-02 -1.27544375e-01 1.63712244e-01 3.36816934e-04
2.26230028e-02 -2.45023393e-05 4.95635588e-03 1.44779564e-01
3.71625177e-02 3.65675948e-03 2.83694017e-02 -7.10500602e-02
-1.15467702e-01 6.21712129e-03 -4.80958959e-02 2.21021066e-02
7.99062499e-02 -1.87164076e-02 -3.67100369e-02 -2.38923731e-02]
[ 6.85871605e-01 5.07725024e-01 8.71024642e-03 3.34823455e-02
4.58684961e-02 9.44384189e-17 -4.46829296e-02 -2.91296778e-02
4.15466461e-02 2.89628330e-02 1.88624017e-03 5.37110446e-02
2.59579053e-03 1.39327071e-02 -2.90781778e-02 5.07209866e-03
5.83403000e-02 2.60764440e-02 4.45999706e-17 -6.66701417e-03
3.03215873e-01 2.26172533e-02 2.43105960e-02 4.98861041e-18
-2.45530791e-02 6.26940708e-02 1.21058073e-02 2.76675948e-04
2.63980996e-02 2.58302364e-02 7.47856723e-02 8.42728943e-02
5.70989097e-02 6.92955086e-02 -5.68313712e-03 1.32199452e-01
8.90511238e-02 -3.45204621e-02 -1.05445836e-01 6.03864150e-03
2.16291384e-02 8.22303162e-03 1.00856715e-02 6.28973151e-02
6.26727169e-02 6.15399206e-02 9.67320897e-02 1.03045269e-16
1.79688783e-01 -1.59960520e-02 -1.15422952e-02 9.60200470e-03
6.58396672e-02 7.78329830e-03 6.53226955e-02 2.45778685e-03
4.36694753e-03 5.75098762e-03 -2.48896201e-02 8.33740755e-05]
Mer drucke di Schaltkreis-Jrööß un di Zwei-Qubit-Gate-Deef uß.
print(f"qubits: {qc.num_qubits}")
print(
f"2q-depth: {transpiled_circuit.depth(lambda x: x.operation.num_qubits==2)}"
)
print(
f"2q-size: {transpiled_circuit.size(lambda x: x.operation.num_qubits==2)}"
)
print(f"Operator counts: {transpiled_circuit.count_ops()}")
transpiled_circuit.draw("mpl", fold=-1, style="clifford", idle_wires=False)
qubits: 60
2q-depth: 64
2q-size: 1888
Operator counts: OrderedDict({'rz': 6016, 'sx': 4576, 'cz': 1888, 'x': 896, 'barrier': 31})

Mer künne jetz över däm jesampte Trainings-Datensatz loope, öm all 1-RDMs ze kräje.
Mer levvere och di Erjebnisse vun enem Experiment, dat mer op Quante-Hardware jelaufe losse han. Ühr künnt entweder dat Training sellef loufe losse, endämm mer dat Flag onge op True setzt, oder di Projektions-Erjebnisse brauche, di mer levvere.
# Set this to True if you want to run the training on hardware
run_experiment = False
# Identity operator on all qubits
id = "I" * num_qubits
# projections_train[i][j][k] will be the expectation value of the j-th Pauli operator (0: X, 1: Y, 2: Z)
# of datapoint i on qubit k
projections_train = []
jobs_train = []
# Experiment options for error mitigation
num_randomizations = 300
shots_per_randomization = 100
noise_factors = [1, 3, 5]
experimental_opts = {}
experimental_opts["resilience"] = {
"measure_mitigation": True,
"zne_mitigation": True,
"zne": {
"noise_factors": noise_factors,
"amplifier": "gate_folding",
"return_all_extrapolated": True,
"return_unextrapolated": True,
"extrapolated_noise_factors": [0] + noise_factors,
},
}
experimental_opts["twirling"] = {
"num_randomizations": num_randomizations,
"shots_per_randomization": shots_per_randomization,
"strategy": "active-accum",
}
options = EstimatorOptions(experimental=experimental_opts)
if run_experiment:
with Batch(backend=backend):
for i in tqdm.tqdm(
range(len(train_data)), desc="Training data progress"
):
# Get training sample
parameters = train_data[i]
# Bind parameter to the circuit and simplify it
qc_bound = qc.assign_parameters(parameters)
transpiler = generate_preset_pass_manager(
optimization_level=3, basis_gates=["u3", "cz"]
)
transpiled_circuit = transpiler.run(qc_bound)
# Transpile for hardware
transpiler = generate_preset_pass_manager(
optimization_level=3, target=target
)
transpiled_circuit = transpiler.run(transpiled_circuit)
# We group all commuting observables
# These groups are the Pauli X, Y and Z operators on individual qubits
observables_x = [
SparsePauliOp(id[:i] + "X" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_y = [
SparsePauliOp(id[:i] + "Y" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_z = [
SparsePauliOp(id[:i] + "Z" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
# We define the primitive unified blocs (PUBs) consisting of the embedding circuit,
# set of observables and the circuit parameters
pub_x = (transpiled_circuit, observables_x)
pub_y = (transpiled_circuit, observables_y)
pub_z = (transpiled_circuit, observables_z)
# We define and run the estimator to obtain <X>, <Y> and <Z> on all qubits
estimator = Estimator(options=options)
job = estimator.run([pub_x, pub_y, pub_z])
jobs_train.append(job)
Training data progress: 100%|██████████| 172/172 [13:03<00:00, 4.55s/it]
Wann di Jobs komplett sin, künne mer di Erjebnisse holle.
if run_experiment:
for i in tqdm.tqdm(
range(len(train_data)), desc="Retrieving training data results"
):
# Completed job
job = jobs_train[i]
# Job results
job_result_x = job.result()[0].data.evs
job_result_y = job.result()[1].data.evs
job_result_z = job.result()[2].data.evs
# Record <X>, <Y> and <Z> on all qubits for the current datapoint
projections_train.append([job_result_x, job_result_y, job_result_z])
Mer widderholle dat för däm Test-Satz.
# Identity operator on all qubits
id = "I" * num_qubits
# projections_test[i][j][k] will be the expectation value of the j-th Pauli operator (0: X, 1: Y, 2: Z)
# of datapoint i on qubit k
projections_test = []
jobs_test = []
# Experiment options for error mitigation
num_randomizations = 300
shots_per_randomization = 100
noise_factors = [1, 3, 5]
experimental_opts = {}
experimental_opts["resilience"] = {
"measure_mitigation": True,
"zne_mitigation": True,
"zne": {
"noise_factors": noise_factors,
"amplifier": "gate_folding",
"return_all_extrapolated": True,
"return_unextrapolated": True,
"extrapolated_noise_factors": [0] + noise_factors,
},
}
experimental_opts["twirling"] = {
"num_randomizations": num_randomizations,
"shots_per_randomization": shots_per_randomization,
"strategy": "active-accum",
}
options = EstimatorOptions(experimental=experimental_opts)
if run_experiment:
with Batch(backend=backend):
for i in tqdm.tqdm(range(len(test_data)), desc="Test data progress"):
# Get test sample
parameters = test_data[i]
# Bind parameter to the circuit and simplify it
qc_bound = qc.assign_parameters(parameters)
transpiler = generate_preset_pass_manager(
optimization_level=3, basis_gates=["u3", "cz"]
)
transpiled_circuit = transpiler.run(qc_bound)
# Transpile for hardware
transpiler = generate_preset_pass_manager(
optimization_level=3, target=target
)
transpiled_circuit = transpiler.run(transpiled_circuit)
# We group all commuting observables
# These groups are the Pauli X, Y and Z operators on individual qubits
observables_x = [
SparsePauliOp(id[:i] + "X" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_y = [
SparsePauliOp(id[:i] + "Y" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_z = [
SparsePauliOp(id[:i] + "Z" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
# We define the primitive unified blocs (PUBs) consisting of the embedding circuit,
# set of observables and the circuit parameters
pub_x = (transpiled_circuit, observables_x)
pub_y = (transpiled_circuit, observables_y)
pub_z = (transpiled_circuit, observables_z)
# We define and run the estimator to obtain <X>, <Y> and <Z> on all qubits
estimator = Estimator(options=options)
job = estimator.run([pub_x, pub_y, pub_z])
jobs_test.append(job)
Test data progress: 100%|██████████| 74/74 [00:13<00:00, 5.56it/s]
Mer künne di Erjebnisse wi vörher holle.
if run_experiment:
for i in tqdm.tqdm(
range(len(test_data)), desc="Retrieving test data results"
):
# Completed job
job = jobs_test[i]
# Job results
job_result_x = job.result()[0].data.evs
job_result_y = job.result()[1].data.evs
job_result_z = job.result()[2].data.evs
# Record <X>, <Y> and <Z> on all qubits for the current datapoint
projections_test.append([job_result_x, job_result_y, job_result_z])
Schrett 4: Nohverarbeidung un Rückjaab em jewünschte klassische Format
Däm projected quantum kernel definiere
Dä projected quantum kernel es definiert met dä folgende Kernel-Funktion:
En dä ovvere Gleichung es e aanpassbare Hyperparameter. Di sin di Einträje vun de Kernel-Matrix .
Mer bruche di Definition vun 1-RDMs, öm ze sinn, dat di einzelne Terme innerhalb vun dä Kernel-Funktion als bewertet wäde künne, wo . Diss Erwartungswerter sin jenaau dat, wat mer ovve jemesse han.
Endämm mer scikit-learn brauche, künne mer däm Kernel söjar noch einvasher berechne. Dat läiet aan dä janz einfach zojängliche radiale Basisfunktion ('rbf') Kernel: . Zeesch mösse mer einfach di neue projizierte Trainings- un Test-Datensätze en zweidimensionale Arrays ömforme.
Luurt druf, dat dat Durchloope vum jesampte Datensatz etwa 80 Minute op dä QPU brauch. Öm secherzestelle, dat dä Räs vum Tutorial einfach usjefööht wäde kann, levvere mer zusätzlich Projektione vun enem vörhär jefaahrete Experiment (di sin en dä Dateie enthalde, di Ühr em Download dataset Code-Block eronjerjelade habt). Wann Ühr dat Training sellef durchjefööht habt, künnt Ühr dat Tutorial met Ühr eijene Erjebnisse forsetze.
if run_experiment:
projections_train = np.array(projections_train).reshape(
len(projections_train), -1
)
projections_test = np.array(projections_test).reshape(
len(projections_test), -1
)
else:
projections_train = np.loadtxt("projections_train.txt")
projections_test = np.loadtxt("projections_test.txt")
Support Vector Machine (SVM)
Mer künne jetz en klassische SVM op dämm vörberechnete Kernel loufe losse, un däm Kernel zwesche Test un Trainings-Sätze för Vörhersooch brauche.
# Range of 'C' and 'gamma' values as SVC hyperparameters
C_range = [0.001, 0.005, 0.007]
C_range.extend([x * 0.01 for x in range(1, 11)])
C_range.extend([x * 0.25 for x in range(1, 60)])
C_range.extend(
[
20,
50,
100,
200,
500,
700,
1000,
1100,
1200,
1300,
1400,
1500,
1700,
2000,
]
)
gamma_range = ["auto", "scale", 0.001, 0.005, 0.007]
gamma_range.extend([x * 0.01 for x in range(1, 11)])
gamma_range.extend([x * 0.25 for x in range(1, 60)])
gamma_range.extend([20, 50, 100])
param_grid = dict(C=C_range, gamma=gamma_range)
# Support vector classifier
svc = SVC(kernel="rbf")
# Define the cross validation
cv = StratifiedKFold(n_splits=10)
# Grid search for hyperparameter tuning (q: quantum)
grid_search_q = GridSearchCV(
svc, param_grid, cv=cv, verbose=1, n_jobs=-1, scoring="f1_weighted"
)
grid_search_q.fit(projections_train, train_labels)
# Best model with best parameters
best_svc_q = grid_search_q.best_estimator_
print(
f"The best parameters are {grid_search_q.best_params_} with a score of {grid_search_q.best_score_:.4f}"
)
# Test accuracy
accuracy_q = best_svc_q.score(projections_test, test_labels)
print(f"Test accuracy with best model: {accuracy_q:.4f}")
Fitting 10 folds for each of 6622 candidates, totalling 66220 fits
The best parameters are {'C': 8.5, 'gamma': 0.01} with a score of 0.6980
Test accuracy with best model: 0.8108
Klassischer Benchmarking
Mer künne en klassische SVM met dä radiale Basisfunktion als Kernel loufe losse, ohne en Quante-Projektion ze maache. Dat Ergebnis es uns klassische Benchmark.
# Support vector classifier
svc = SVC(kernel="rbf")
# Grid search for hyperparameter tuning (c: classical)
grid_search_c = GridSearchCV(
svc, param_grid, cv=cv, verbose=1, n_jobs=-1, scoring="f1_weighted"
)
grid_search_c.fit(train_data, train_labels)
# Best model with best parameters
best_svc_c = grid_search_c.best_estimator_
print(
f"The best parameters are {grid_search_c.best_params_} with a score of {grid_search_c.best_score_:.4f}"
)
# Test accuracy
accuracy_c = best_svc_c.score(test_data, test_labels)
print(f"Test accuracy with best model: {accuracy_c:.4f}")
Fitting 10 folds for each of 6622 candidates, totalling 66220 fits
The best parameters are {'C': 10.75, 'gamma': 0.04} with a score of 0.7830
Test accuracy with best model: 0.7432
Aanhang: Vörsicht — däm Datensatz sing möjliche Quantevördeil för Lernufjaabe prööve
Nit all Datensätze beede möjliche Vördeile dörch di Nutzung vun PQKs. Et jitt theoretische Jränze, di mer als vörläufije Test brauche kann, öm ze luure, ov en bestemmpte Datensatz vun PQKs profitiere kann. Öm dat ze quantifiziere, definiere di Autore vun Power of data in quantum machine learning [2] Jrößte, di als klassische un Quante-Modellkomplexitäte un jeometrische Trennung vun dä klassische un Quante-Modelle bezeechnet wäde. Öm ene möjliche Quantevördeil vun PQKs ze erwarte, sull di jeometrische Trennung zwesche dä klassische un Quante-projizierte Kernels ungefähr en dä Jevend vun ligge, wo di Aanzahl vun Trainings-Samples es. Wann diss Bedingung erfüllt es, maache mer wigger met däm Prööfe vun dä Modellkomplexitäte. Wann di klassische Modellkomplexität en dä Jevend vun läiet, während di Quante-projizierte Modellkomplexität wesentlich kleiner als es, künne mer ene möjliche Vördeil vum PQK erwarte. Jeometrische Trennung es wi folgk definiert (F19 en [2]):
# Gamma values used in best models above
gamma_c = grid_search_c.best_params_["gamma"]
gamma_q = grid_search_q.best_params_["gamma"]
# Regularization parameter used in the best classical model above
C_c = grid_search_c.best_params_["C"]
l_c = 1 / C_c
# Classical and quantum kernels used above
K_c = rbf_kernel(train_data, train_data, gamma=gamma_c)
K_q = rbf_kernel(projections_train, projections_train, gamma=gamma_q)
# Intermediate matrices in the equation
K_c_sqrt = sqrtm(K_c)
K_q_sqrt = sqrtm(K_q)
K_c_inv = inv(K_c + l_c * np.eye(K_c.shape[0]))
K_multiplication = (
K_q_sqrt @ K_c_sqrt @ K_c_inv @ K_c_inv @ K_c_sqrt @ K_q_sqrt
)
# Geometric separation
norm = np.linalg.norm(K_multiplication, ord=np.inf)
g_cq = np.sqrt(norm)
print(
f"Geometric separation between classical and quantum kernels is {g_cq:.4f}"
)
print(np.sqrt(len(train_data)))
Geometric separation between classical and quantum kernels is 1.5440
13.114877048604
Modellkomplexität es wi folgk definiert (M1 en [2]):
# Model complexity of the classical kernel
# Number of training data
N = len(train_data)
# Predicted labels
pred_labels = best_svc_c.predict(train_data)
pred_matrix = np.outer(pred_labels, pred_labels)
# Intermediate terms
K_c_inv = inv(K_c + l_c * np.eye(K_c.shape[0]))
# First term
first_sum = np.sum((K_c_inv @ K_c_inv) * pred_matrix)
first_term = l_c * np.sqrt(first_sum / N)
# Second term
second_sum = np.sum((K_c_inv @ K_c @ K_c_inv) * pred_matrix)
second_term = np.sqrt(second_sum / N)
# Model complexity
s_c = first_term + second_term
print(f"Classical model complexity is {s_c:.4f}")
Classical model complexity is 1.3578
# Model complexity of the projected quantum kernel
# Number of training data
N = len(projections_train)
# Predicted labels
pred_labels = best_svc_q.predict(projections_train)
pred_matrix = np.outer(pred_labels, pred_labels)
# Regularization parameter used in the best classical model above
C_q = grid_search_q.best_params_["C"]
l_q = 1 / C_q
# Intermediate terms
K_q_inv = inv(K_q + l_q * np.eye(K_q.shape[0]))
# First term
first_sum = np.sum((K_q_inv @ K_q_inv) * pred_matrix)
first_term = l_q * np.sqrt(first_sum / N)
# Second term
second_sum = np.sum((K_q_inv @ K_q @ K_q_inv) * pred_matrix)
second_term = np.sqrt(second_sum / N)
# Model complexity
s_q = first_term + second_term
print(f"Quantum model complexity is {s_q:.4f}")
Quantum model complexity is 1.5806
Referenze
- Utro, Filippo, et al. "Enhanced Prediction of CAR T-Cell Cytotoxicity with Quantum-Kernel Methods." arXiv preprint arXiv:2507.22710 (2025).
- Huang, Hsin-Yuan, et al. "Power of data in quantum machine learning." Nature communications 12.1 (2021): 2631.
- Daniels, Kyle G., et al. "Decoding CAR T cell phenotype using combinatorial signaling motif libraries and machine learning." Science 378.6625 (2022): 1194-1200.